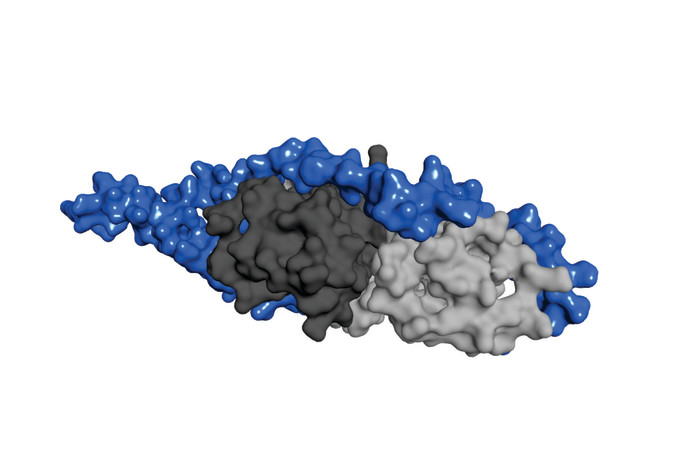
Protein interaction helps Yersinia cause disease
Researchers at Umeå University, Sweden, in collaboration with an international team, have discovered a new mechanism for interaction between two proteins that are vital for the Yersinia pseudotuberculosis bacteria’s pathogenic ability.
A common strategy bacteria have to cause disease is to transfer toxic proteins to host cells, for example in humans. Yersinia pseudotuberculosis is a bacteria that can cause gastrointestinal infections. The bacteria transfers proteins called Yop (Yersinia outer protein) through a complex needle structure forming a pore in the host cell membrane. Yop proteins are made up of segments with different functions. The YopH protein has a segment which counteracts the immun system of the host cells. Another segment binds to chaperones, a group of proteins that help other proteins uphold a correct structure, which is important for transporting YopH through the needle structure to the host cell.
A team of researchers led by professor Magnus Wolf-Watz at the Department of Chemistry, Umeå University, has now discovered how the chaperone binding part of YopH recognizes and interacts with the protein SycH. SycH is a chaperone whose task is to enable YopH to be transported through the needle structure and into the host cell. The research team has discovered that the chaperone binding part of YopH must completely loose its three dimensional structure to be able to grasp around the SycH protein like a horse’s shoe.
”This type of mechanism for protein-protein interaction can be called ”coupled folding and binding” and has not been seen before. Through this discovery we have contributed to a basic understanding of protein-protein interactions” says Magnus Wolf-Watz. ”This is important because many functions inside cells are carried out by protein-protein complexes.”
The discovery became possibly by Magnus Wolf-Watz putting together a team of researchers from different countries and with different special competence. The team consisted of group leaders Anders Hofer of Umeå University (expert in determining stoichiometry in protein complexes), Alexander Schug, Karlsruhe Institute of Technology (expert in modelling protein structures), Dmitri Svergun, EMBL, Hamburg (expert in protein structure determination with SAXS methodology) and Andrew Baldwin, Oxford University (expert in measuring relaxation with NMR spectroscopy). Experimentally the study was led by Arun Gupta, former post doc in Magnus Wolf-Watz’s research group.
Wissenschaftlicher Ansprechpartner:
For more information, please contact:
Magnus Wolf-Watz, professor, Department of Chemistry, Umeå University
Phone: +56907867690
Email: magnus.wolf-watz@umu.se
Originalpublikation:
Original article:
Gupta, A., Reinartz, I., Karunanithy, G., Spilotros, A., Jonna, V.R.,Hofer, A., Svergun, D., Baldwin, A., Schug, A., and Wolf-Watz, M, Formation of a secretion competent protein complex by a dynamic wrap-around binding mechanism, Journal of Molecular Biology, Volume 430, Issue 18, Part B, 14 September 2018, DOI 10.1016/j.jmb.2018.07.014
Weitere Informationen:
https://www.sciencedirect.com/science/article/pii/S002228361830442X
https://mediabank.umu.se/selection/44f28f42e19e8f2ce6272054e17bbb77
Die semantisch ähnlichsten Pressemitteilungen im idw