Additional Genetic Cause for Non-alcoholic Fatty Liver Disease Discovered
In Germany about 18 million people suffer from non-alcoholic fatty liver. The causes of this disease are manifold and include environmental as well as genetic factors. DZD researchers have now discovered new genes that play a role in the development of fatty liver. In humans and mice, respectively, the genes IRGM, Ifgga2 and Ifgga4 are responsible for the production of regulatory proteins of the family of immunity-related GTPases which counteract fat accumulation in the liver. However, a genetic variation leads to the formation of fewer of these proteins. The results have now been published in the Journal of Hepatology.
Non-alcoholic fatty liver disease (NAFLD) is the leading cause of chronic liver disease in Europe and the United States. In Europe about 20-30 percent of the population are affected. NAFLD is often associated with other diseases such as obesity, type 2 diabetes, high blood pressure (arterial hypertension) and a fat metabolism disorder (dyslipidemia). In addition to an unhealthy lifestyle with a high-fat, high-sugar diet and lack of exercise, a genetic predisposition is also responsible for the development of this liver disease. However, for this complex disease not only one gene butrather the interactions of different genes and epigenetic * factors are responsible. Researchers have now discovered a new family of genes that play an important role in preventing fatty liver development. In humans and mice, these genes produce regulatory proteins from the family of immunity-related GTPases that counteract fat accumulation in the liver. However, if there is a genetic modification, fewer proteins are formed. Studies show that the liver of patients with NAFLD and mice with fatty liver have significantly lower amounts of these proteins. The study, which has now been published in the Journal of Hepatology, was carried out by a team of researchers of the German Institute of Human Nutrition Potsdam-Rehbrücke (DIfE), the German Diabetes Center (DDZ) and Helmholtz Zentrum München – all partners of the German Center for Diabetes Research (DZD).
New genes identified
Using molecular markers and statistical methods – quantitative trait locus (QTL) analysis – genes that cause complex human diseases can be identified in mouse strains. For example, the research team discovered a region on mouse chromosome 18 that was associated with altered amounts of fat in the liver. If the genes Ifgga2 and Ifgga4 are expressed, proteins of the family of immunity-related GTPases are formed – in the mouse the proteins IFGGA2 and IFGGA4 and in humans the protein IRGM. These proteins increase a certain form of fat degradation and thus counteract the development of fatty liver.
The reason for a lower expression in mice with a fatty liver is a small genetic variation. "Due to the loss of only a single base in a gene sequence, which increases the expression of a certain gene, the two related proteins IFGGA2 and IFGGA4 are hardly produced in liver cells of mice that are susceptible to fatty liver," said Professor Annette Schürmann, head of the Department of Experimental Diabetology at the German Institute of Human Nutrition Potsdam-Rehbrücke (DIfE) and spokesperson for the German Center for Diabetes Research (DZD). Patients with NAFLD also have significantly lower amounts of the corresponding protein (IRGM). As a result, the fat content in the liver can increase three to four-fold.
Proteins increase specific degradation of fat in the liver (lipophagy)
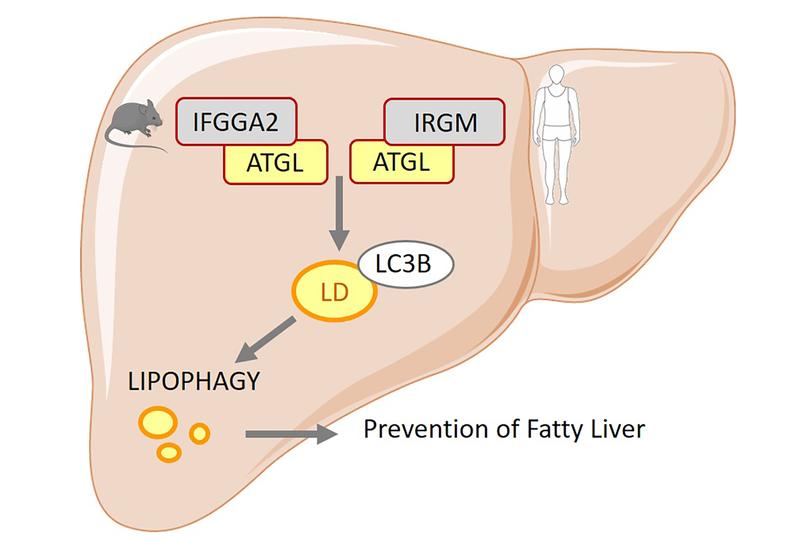
Functional studies have shown that an overproduction of immunity-related GTPases in liver cells or in the liver of mice, significantly reduced their fat content. "The reason for this is the induction of a particular form of autophagy that is specific for the degradation of fats and is therefore called lipophagy," explained Dr. Wenke Jonas, who co-directed the study together with Professor Schürmann. Autophagy is a type of cellular disposal and recycling process through which the cell's own components are degraded. The researchers have observed that after the uptake of fatty acids in liver cells, the immunity-related GTPases migrate to the lipid droplets. There they bind to an enzyme involved in fat degradation (adipocyte triglyceride lipase, ATGL) and ensure that a central protein of autophagy (LC3B) binds to the fat droplet. Due to the autophagy of lipid droplets, the amount of fat is reduced and thus the development of fatty liver is prevented.
The researchers were also able to show that the immunity-related GTPases affect the amount of fat in the liver in the following two studies: If they inhibited the synthesis of the proteins, mice stored more fat in the liver cells. If, on the other hand, the production of the proteins in liver cells was increased, the cells stored considerably less fat.
“Our work has identified further important genes that cause fatty liver disease. The study results also deepen our understanding of which cellular processes have to be stimulated to counteract fatty liver development,” said Schürmann. "Our next goal is to clarify which measures – such as diets or certain drugs – can increase the amount of immunity-related GTPases in order to reduce fat storage in the liver."
* Epigenetics investigates those properties of genes that are not revealed by the DNA sequence itself, but by their expression. Epigenetic information is mediated by methyl groups or other biomolecules which, like chemical locks, deny or release access to certain DNA sequences and thus control their activation. Which epigenetic code is established in a person and whether it changes in the course of life is determined not only by the body's own signal substances but also by eating habits and other aspects of lifestyle.
Wissenschaftlicher Ansprechpartner:
Prof. Dr. Annette Schürmann
Head, Department of Experimental Diabetology
German Institute of Human Nutrition Potsdam-Rehbrücke (DIfE)
Phone: +49 (0)33200 88-2368
email: schuermann(at)dife.de
Originalpublikation:
Schwerbel, K. et al: Immunity-related GTPase induces lipophagy to prevent excess hepatic lipid accumulation. Journal of Hepatology (2020); DOI: https://doi.org/10.1016/j.jhep.2020.04.031
Die semantisch ähnlichsten Pressemitteilungen im idw