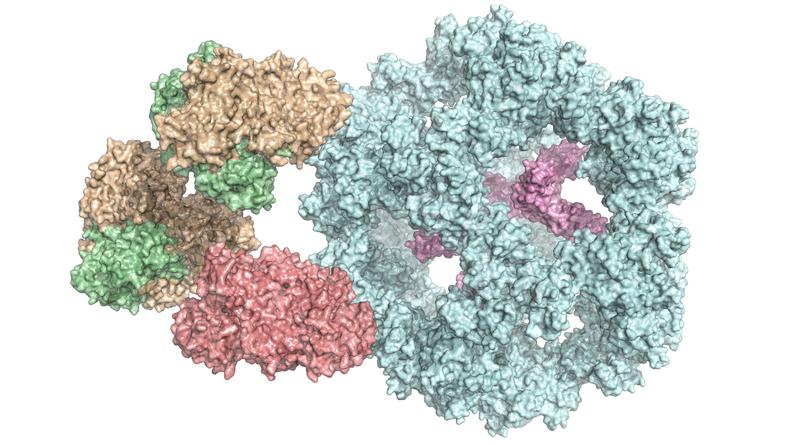
Metabolism: Researchers first to shed light on structure of huge enzyme complex
A new method has enabled the natural structure of particularly large and complex enzymes to be revealed. Scientists at Martin Luther University Halle-Wittenberg (MLU) and TU Berlin have published their findings in the journal "Cell Reports". They investigated a multi-enzyme complex that plays an essential role in metabolism and have discovered that it functions differently than previously thought. This will help scientists better understand certain diseases.
Enzymes are a cell’s biocatalysts. They accelerate chemical reactions in the body or ensure that these reactions even take place at all. As a result, they play an extremely important role in metabolism. Individual enzymes frequently form a complex with many subunits, as in the case of the pyruvate dehydrogenase complex. It occurs in all human, animal and plant cells, as well as in fungi and even in some bacteria. "It is vital for energy production in cells," explains Dr Panagiotis Kastritis, an assistant professor at MLU and group leader at the Centre for Innovation Competence HALOmem.
"Up until now, the enzyme complex has always been broken down into its individual parts before being examined. Studying the entire, functional complex is very complicated," says Kastritis. To solve this problem, his team combined various biochemical and biophysical methods of investigation. In addition to using mass spectrometry and chemical analyses, a special form of electron microscopy - cryo-electron microscopy - has been essential for studying such a complex structure, says the structural biologist. It enables complex, organic structures to be observed at a high resolution and in their natural state. "However, it is very important to subsequently bring together all this data in computer models," says Kastritis. For the current study, the team of researchers is using cell extracts of a fungus that can survive at high temperatures, which makes analysis easier.
The scientists have thus gained an insight into how the various subunits of the enzyme complex interact with each other and have also discovered that it functions differently than previously assumed. "It was previously thought that all of the subunits were directly involved in the reaction," says Kastritis. "But now we know that some of them seem to form a kind of chamber that protects the reaction." Because the complexes are very similar across organisms, the results are also helping researchers understand the human pyruvate dehydrogenase complex.
The new method could help scientists better understand many different diseases. Some viruses, for example, reduce the number of enzyme associations in the cells. This effect has also been observed in Alzheimer’s disease. In some cases, however, the enzymes do not function properly and the vital reactions occur either too slowly or too quickly. "If we are able to better understand the natural structure, we will be able to recognise tiny changes and understand why the reaction is no longer occurring as it should," says Kastritis. This knowledge could theoretically also form the basis for new treatment options.
The study was funded by the Federal Ministry of Education and Research (BMBF), the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation), the Wellcome Trust and with funds from the European Regional Development Fund (ERDF).
Wissenschaftlicher Ansprechpartner:
Assistant Professor Panagiotis Kastritis
Institute of Biochemistry and Biotechnology / MLU
ZIK HALOmem "membrane structures and dynamics"
phone: +49 345 55-24983
email: panagiotis.kastritis@bct.uni-halle.de
web: https://blogs.urz.uni-halle.de/kastritislab/
Originalpublikation:
Kyrilis F. L. et al. Integrative structure of a 10-megadalton eukaryotic pyruvate dehydrogenase complex from native cell extracts. Cell Reports (2021). https://doi.org/10.1016/j.celrep.2021.108727
Ähnliche Pressemitteilungen im idw