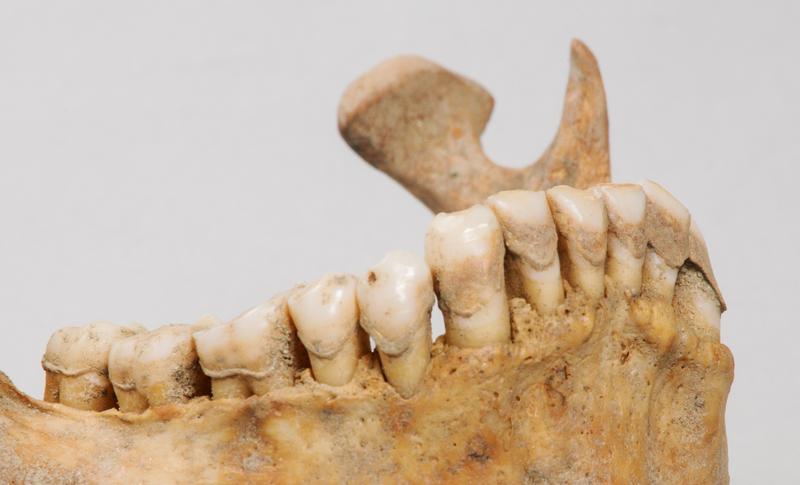
Mummy research: Ancient dental calculus - new insights into the evolution of oral microbiota
A research team from Eurac Research and the University of Trento has examined ancient calculus samples from skeletal remains from South Tyrol and Trentino and discovered previously unknown species of microorganisms
Plaque – horrible stuff, right? Well yes, but not entirely: old tooth plaque also known as dental calculus samples provide a valuable source of information about our oral microbiota and its development. A team of researchers from Eurac Research and the University of Trento, examined the calculus of twenty human skeletal remains from South Tyrol and Trentino dating from the Neolithic period to the Early Middle Ages. The team discovered two previously unknown species of a common microorganism in our bodies called Methanobrevibacter. Thanks to the application of a bioinformatics method not yet established in mummy research, the research group was also able to reveal that on a temporal axis of 50,000 years, the diversity of this organism in our oral flora has declined sharply in recent centuries. The results of the study have now been published in the renowned journal "Microbiome".
The remains in question range from about 5,500 to 1,000 years old and were discovered at several burial sites in Trentino and South Tyrol. Of the 20 individuals examined, 18 exhibited oral diseases, especially periodontitis, a bacterial inflammation of the gums. In the past, people didn’t brush their teeth like they do today and as a result, deposits of large amounts of tartar with well-preserved DNA have survived the archaeological record. Following an initial analysis, the research group focused primarily on the main representative that was present in abundance and in great diversity among the microorganisms in the tartar - Methanobrevibacter, an archaeal microorganism, common in the human digestive tract.
In order to investigate the diversity of this genus, the team decided on a new methodological approach, namely, to decode the entire genomes of the existing variants of Methanobrevibacter. In addition to the skeletal remains, the researchers also worked through 82 previously published calculus studies - including one from a Neanderthal find - analyzing in total 102 calculus samples spanning a period of over 50,000 years, eight countries and three continents.
"It was the first time we applied the method so consistently and with such breadth, and we didn't know what we would come across," explains microbiologist Lena Granehäll, lead author of the study. "It was an 'aha' moment when we then realised that we had discovered two completely new species of Methanobrevibacter and were thus also able to reveal a genetic diversity of this microorganism that was previously unknown. No geographical patterns are discernible here; all species occurred in the samples in Europe as well as Asia and America."
However, one thing was discernable. The research team observed a clear tendency in terms of time: while the newly discovered species turned out to be "older" representatives of Methanobrevibacter and were consistently present in the oldest tartar samples up to the Late Middle Ages, they were replaced by a single species from that time onwards. In the samples from the Middle Ages onwards, the older representatives are completely absent. "As our study shows, there seems to have been a loss of species of this microorganism in our mouths. It goes hand in hand with what we have also already seen in stomach samples, namely, 5,000 years ago, there was a diversity of bacteria that decreased significantly over a very short period of time," underlines Eurac Research microbiologist Frank Maixner. "Genetically, few changes have occurred in humans over the last 5,000 years; but the same cannot be said for our commensal bacteria which have been exposed to greater influences: we see that industrialisation, new therapies, changes in medicine and in our diet, have all impacted the composition of our bacterial diversity, not only in the mouth, but all the way to the stomach and intestines too."
Nicola Segata, who works at the Department of Cellular, Computational, and Integrative Biology (CIBIO) at the University of Trento specialises in the study of human microbiome -the totality of microorganisms in and on our bodies. During the course of dedicated years of research, he has catalogued the microorganisms of the intestinal microbiome and in doing so, also discovered many species of bacteria that were previously unknown.
For the calculus study, Segata and his team sequenced the DNA that had been extracted from the ancient samples by the Eurac Research team. "An important part of the work in this study was to separate old DNA from modern DNA after sequencing the genetic material. Because old samples like these remained in a non-sterile environment for many many years, they contain a lot of microbes and DNA which do not belong to the original oral microbiome," explains Nicola Segata. In his laboratory in Trento, he researches both modern and older samples to learn more about the human microbiome. "Analysing old samples is a bit like studying genetic fossils and in this way, we can reconstruct the evolution of particular microorganisms. As in this case, we can also discover new species of microorganisms that we may have lost over time due to our modern lifestyle," Segata concludes.
Originalpublikation:
https://doi.org/10.1186/s40168-021-01132-8
Ähnliche Pressemitteilungen im idw